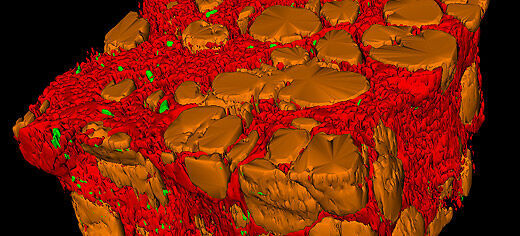
Computing experts and medical researchers at the University of Leeds have developed a fast, easy-to-use way of studying tissue samples in 3D using 'virtual' microscope slides.
The novel digital scanning system produces high-resolution, multicoloured images that can be rotated and examined from any angle.
The new approach is revealing more information about disease processes - information that could be used in future to develop new therapies or explain why conventional treatments are not working. Such 3D views of tissue samples may also eventually play a role in clinical practice, as medical imaging technology provides even higher resolution images of tissue.
Digital 3D reconstruction of tissue has many uses in biological and medical research. Viewing tissue in 3D allows researchers to study its shape in ways that would not be possible with conventional methods. For example, a biologist may want to study the structure of developing organs, a cancer specialist may study the branching of blood vessels supplying a tumour, or a liver specialist may need to understand how this vital organ reacts to damage caused by hepatitis C. All of these require an understanding of the shape of the tissue in three dimensions.
At the moment, hospital pathologists and medical researchers cut tissue samples into ultra-thin slices and examine these by hand, one-by-one, on a microscope. This is a fairly labour-intensive process - a single slide can contain several hundred thousand cells - and the number of slices examined will be limited by the time available. To do a true 3D analysis, users would need to look at 100s of different 2D sections - something that would be prohibitively expensive by hand.
In contrast, the system developed at the University of Leeds requires almost no extra manual input once the tissue has been cut and mounted onto glass slides. An automated system turns batches of the slides into high-resolution digital images, which are then aligned using image registration software. Users can then study these virtual blocks of tissue in 3D and zoom in on particular areas of interest.
The researchers have now tested the system on eight different types of tissue, using more than 13000 virtual slides to create around 400 separate 3D volumes. The system and selected case studies, including examples of liver disease, cancer and embryology, are described in the May issue of the American Journal of Pathology.
"Up until now, the use of 3D imaging technology to study disease has been limited because of low resolution, and the time and difficulty associated with acquiring large numbers of images with a microscope," said lead investigator Dr. Darren Treanor, pathologist at the University of Leeds and the Leeds Teaching Hospitals NHS Trust. "Our virtual system means that users can look at the shape and structure of cells and the 'micro-architecture' of blood vessels and tumours on large tissue samples. This can all be done without input from computing specialists."
"Having a 3D view can often make a real difference," said Dr Derek Magee, from the University of Leeds' School of Computing who developed the software behind the system. "For instance, if you want to understand how a system of blood vessels supplying a tumour connects up, you really need to see that in 3D, not as a series of separate 2D sections."
The work was funded by the National Cancer Research Institute informatics initiative, Leeds Teaching Hospital Trust Research and Development, National Institute for Health Research, West Yorkshire Comprehensive Local Research Network, UK Department of Health, and the Wellcome Trust.
Further information:
Please contact the University of Leeds Press Office on +44 (0)113 343 4031 or email pressoffice@leeds.ac.uk
Examples of the 3D images and the virtual slides used to create them can be found at http://www.virtualpathology.leeds.ac.uk/.
"Towards Routine Use of 3D Histopathology As a Research Tool," by N. Roberts, D. Magee, Y. Song, K. Brabazon, M. Shires, D. Crellin, N.M. Orsi, R. Quirke, P. Quirke, and D. Treanor (DOI: http://dx.doi.org/10.1016/j.ajpath.2012.01.033) will appear in the American Journal of Pathology, Volume 180, Issue 5 (May 2012) published by Elsevier.