Hospital screening tests are failing to identify the true extent of microbial resistance, according to new research.
Scientists have found that disease-causing bacteria carry antibiotic resistance genes that are dormant as a result of a genetic mutation. During screening, the bacteria appear susceptible to antibiotics.
Our findings highlight the fact that antibiotic resistance can sometimes effectively be hidden ... it looks like you're dealing with an infection that should be treatable but that may not be the case.
But such mutations easily become lost, rapidly transforming the bacteria into drug-resistant strains – dramatically increasing the risk that the drug therapy will fail.
The study, led by the University of Leeds, reveals another layer in the complex scientific fight to understand and combat one of the biggest challenges facing modern healthcare: the rise of drug-resistant superbugs.
Dr Alex O’Neill, Associate Professor in the School of Molecular and Cellular Biology and Astbury Centre for Structural Molecular Biology at Leeds, said: “Our findings highlight the fact that antibiotic resistance can sometimes effectively be hidden. In other words, it looks like you’re dealing with an infection that should be treatable with a given antibiotic, but that may not be the case.”
The researchers analysed nearly 1,500 samples of Staphylococcus aureus – a bacteria that is commonly found on the skin but can cause more serious infections and toxic-shock syndrome. It is best known in its highly drug-resistant form, MRSA, which frequently causes a potentially-lethal hospital-acquired infection.
In the study, published in the journal mBio, the scientists refer to the process by which the drug resistance gene is de-activated as SARM – Silencing of Antibiotic Resistance by Mutation.
About three per cent of the samples had the silenced resistance gene and were tested as being susceptible to antibiotic treatment.
But in nine out of ten samples where SARM was identified, the effect of the mutation was quickly lost when the bacteria was exposed to antibiotics.
Antibiotic resistance is rapidly restored
This was due to a process called "reversion" – in effect a second mutation which restored the pathogen to its original, drug-resistant state.
Dr O’Neill said: “The ease with which SARM strains become drug-resistant means that they are ‘wolves in sheep’s clothing’. SARM is therefore a potentially important reason why antibiotic treatments fail unexpectedly in infected patients.
“We hope that our study will alert doctors to the knowledge that silenced antibiotic resistance genes are relatively common in bacteria such as S. aureus, and therefore have significant potential to adversely affect antibiotic treatment.”
The research reveals a limitation in the routine screening process for drug-resistant bacteria.
Bacteria isolated from patients with a serious infection undergo what is known as susceptibility testing – tests to determine which antibacterial drugs will be effective in attacking the infection. This is the gold-standard approach but the research reveals how the test is "blindsided" in SARM strains where the resistance gene is dormant.
Dr O’Neill added: “SARM can be detected using DNA sequencing to determine the complete genetic make-up of strains, but this technology is not yet in routine use in clinical microbiology laboratories.”
Although the study only looked at S. aureus, the researchers believe it is likely that this process also occurs on other bacteria.
Challenge of superbugs
The NHS says the overuse of antibiotics worldwide has accelerated the development of drug resistant pathogens, the so called superbugs.
The University of Leeds is part of an international fight to tackle the threat of antimicrobial resistance with research focusing on a number of fronts, from understanding the molecular biology of pathogens, to better ways of identifying bacterial infections and raising public awareness of the need for rational antibiotic use.
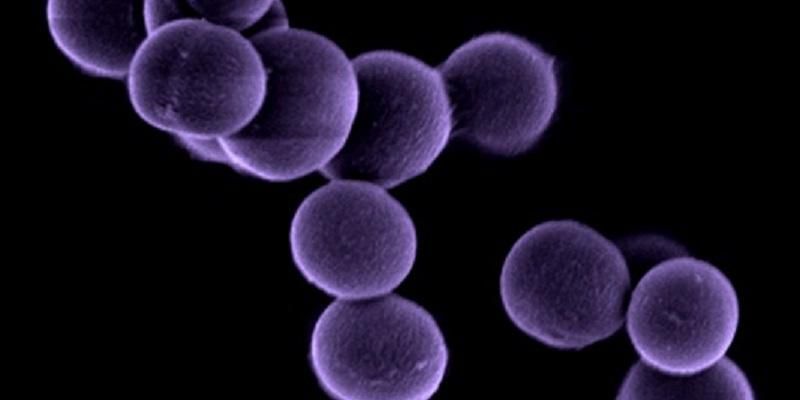
Image: A picture taken with an electron microscope showing the Staphylococcus aureus bacteria.