Leeds is leading pioneering research into new technology for imaging babies with congenital heart disease (CHD).
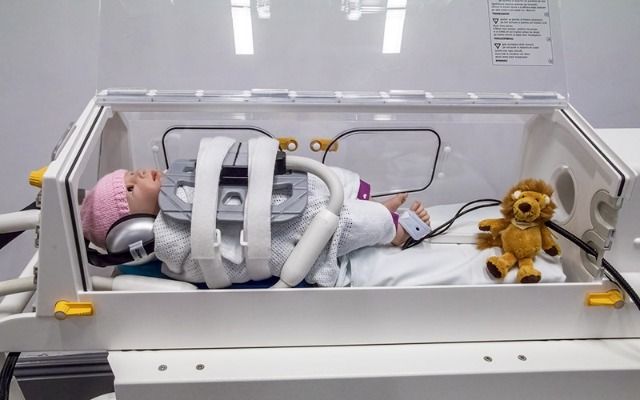
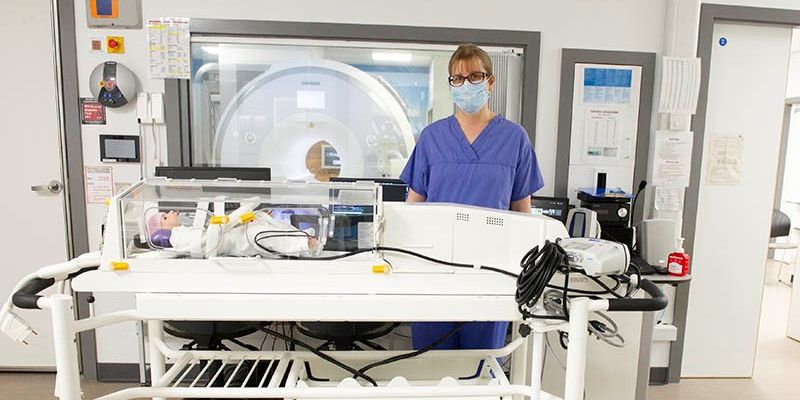
The Baby MRI incubator at Leeds Congenital Heart Unit (LCHU), currently being trialled by researchers in the School of Medicine, is the first of its kind in the UK. It is part of a suite of state-of-the-art equipment providing detailed heart imaging during MRI scans.
Leading clinicians say the improved data may give worried parents an earlier diagnosis for their poorly babies, and help predict which patients need surgery in the future.
The research we are currently undertaking makes the Leeds Congenital Heart Unit the first in the UK to scan babies with congenital heart disease this way.
It was funded by Children's Heart Surgery Fund, with supporting donations from Morrisons Foundation, Heart Research UK, Ilkley Round Table and other key funders.
The incubator allows doctors to use the latest 4D Flow MRI technology which will not only reduce diagnostic uncertainty, but also lower the risks associated with traditional imaging methods and the scan time to under 10 minutes.
Dr Malenka Bissell, Clinical Lecturer of Paediatric Cardiology in the School of Medicine, said: “The research we are currently undertaking makes the Leeds Congenital Heart Unit the first in the UK to scan babies with congenital heart disease this way.
“In combination with the equipment funded by the charity, 4D Flow MRI technology is paving the way to improve diagnostic pathways for these patients.
“Uncertainty can be one of the most stressful parts for parents, so we're now looking at how we can use advanced imaging markers to improve our ability to predict when and if a patient will need surgery.”
According to the NHS, congenital heart disease is one of the most common types of birth defect, affecting up to eight in every 1,000 babies born in the UK.
The challenge of diagnosis
While survival rates have improved, diagnosis and imaging of newborns can be challenging.
Traditionally, CT imaging has often been preferred over MRI, due to its quieter and faster scan times, while MRI has necessitated breath holds and patient sedation with its associated risk to babies.
The use of CT however requires a radiation dose and produces less detailed data than an MRI.
The new incubator will also allow babies to be transported directly from the ward into the MRI department, where scans can be conducted free from complications and distress.
Game-changing research
Children's Heart Surgery Fund CEO Sharon Milner said: "I must pay tribute to the many trusts, foundations and other funders who helped us purchase this vital technology - thank you for supporting hearts for life.
“This game-changing research will make a huge difference to babies born with heart disease and their families for years to come in our region.”
It is understood thousands of babies will benefit from research studies into paediatric congenital heart disease incorporating the Baby MRI Incubator over the coming years.
Leeds' heart surgery success
The MRI Baby Pod research is happening in the wake of a hugely positive December 2020 report from the National Institute for Cardiovascular Outcomes Research (NICOR), showing children who undergo heart surgery in Leeds have better outcomes than anywhere else in the country.
Over a three-year period, Leeds Congenital Heart Unit performed heart surgery on 929 children and achieved a survival rate of 99.4 per cent.
Leeds was the only children's heart surgery unit in the UK to have a survival rate that was in the highest performance category, being much higher than expected for units of its kind in England.
Children's Heart Surgery Fund have supported the LCHU for over 30 years, since its formation in 1988.
Further information
Main image: The baby pod. Credit: Malenka Bissell
For media enquiries email University of Leeds Press Officer Lauren Ballinger at l.ballinger@leeds.ac.uk.